- Denaturation at 94°C :
- Annealing at 54°C :
- extension at 72°C :
During the denaturation, the double strand melts open to single stranded DNA, all enzymatic reactions stop (for example : the extension from a previous cycle).
The primers are jiggling around, caused by the Brownian motion. Ionic bonds are constantly formed and broken between the single stranded primer and the single stranded template. The more stable bonds last a little bit longer (primers that fit exactly) and on that little piece of double stranded DNA (template and primer), the polymerase can attach and starts copying the template. Once there are a few bases built in, the ionic bond is so strong between the template and the primer, that it does not break anymore.
This is the ideal working temperature for the polymerase. The primers, where there are a few bases built in, already have a stronger ionic attraction to the template than the forces breaking these attractions. Primers that are on positions with no exact match, get loose again (because of the higher temperature) and don't give an extension of the fragment.
The bases (complementary to the template) are coupled to the primer on the 3' side (the polymerase adds dNTP's from 5' to 3', reading the template from 3' to 5' side, bases are added complementary to the template)
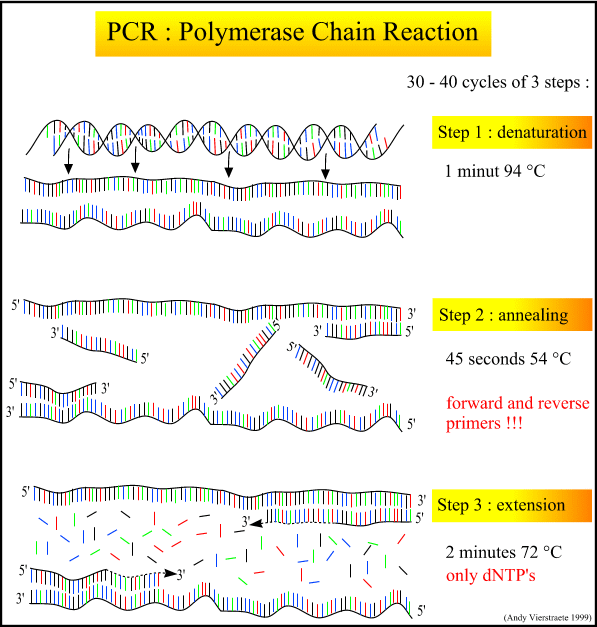
Figure 3 : The different steps in PCR. (pdf file of this picture) Animated picture of PCR Because both strands are copied during PCR, there is an exponential increase of the number of copies of the gene. Suppose there is only one copy of the wanted gene before the cycling starts, after one cycle, there will be 2 copies, after two cycles, there will be 4 copies, three cycles will result in 8 copies and so on.
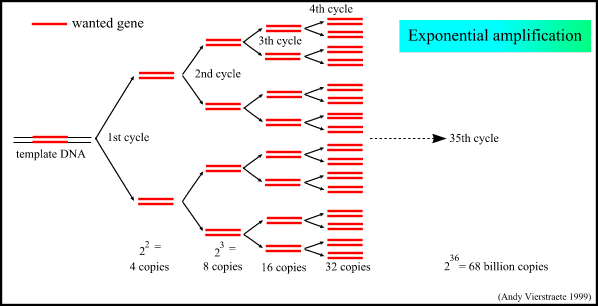
Figure 4 : The exponential amplification of the gene in PCR.
- There is a product formed.
Though biochemistry is an exact science, not every PCR is successful. There is for example a possibility that the quality of the DNA is poor, that one of the primers doesn't fit, or that there is too much starting template - The product is of the right size
It is possible that there is a product, for example a band of 500 bases, but the expected gene should be 1800 bases long. In that case, one of the primers probably fits on a part of the gene closer to the other primer. It is also possible that both primers fit on a totally different gene. - Only one band is formed.
As in the description above, it is possible that the primers fit on the desired locations, and also on other locations. In that case, you can have different bands in one lane on a gel.
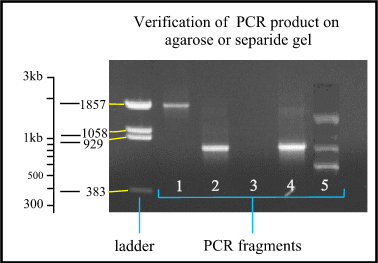
Figure 5 : Verification of the PCR product on gel. The ladder is a mixture of fragments with known size to compare with the PCR fragments. Notice that the distance between the different fragments of the ladder is logarithmic. Lane 1 : PCR fragment is approximately 1850 bases long. Lane 2 and 4 : the fragments are approximately 800 bases long. Lane 3 : no product is formed, so the PCR failed. Lane 5 : multiple bands are formed because one of the primers fits on different places.